The global autoimmune disorder therapies market was valued at $53.2 billion in 2019. The market is expected to grow at a compound annual growth rate (CAGR) of 11.2% to reach $90.7 billion by 2024.
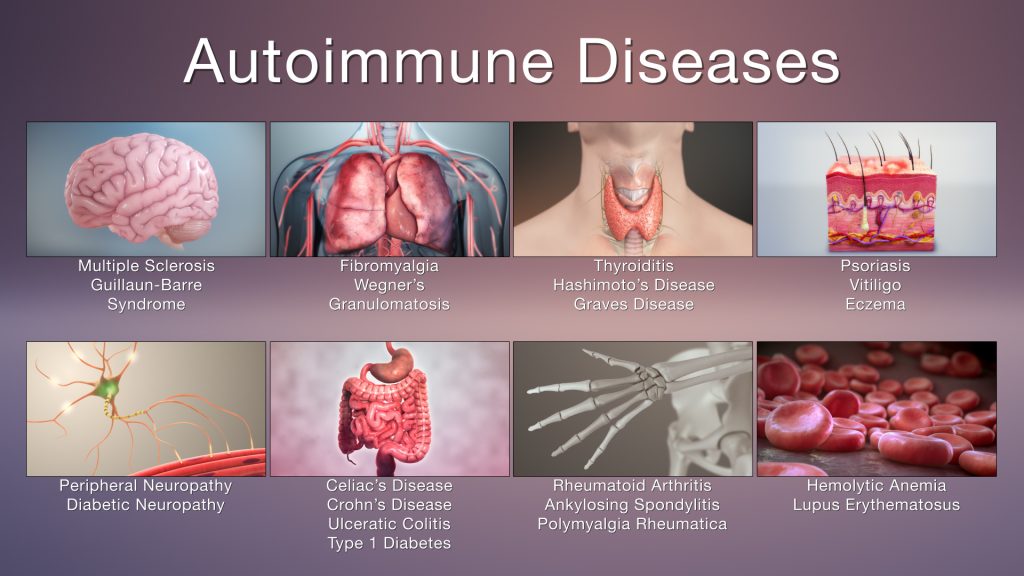
Growth of the global market is attributed to factors such as the growing prevalence of obesity, type 1 diabetes and other chronic diseases which leads to autoimmune diseases, a strong product regulatory scenario, and strong investment in research and development activities by key market players including AstraZeneca, Vertex Pharmaceuticals Incorporated, Bristol-Myers Squibb, F. Hoffmann-La Roche AG (Roche), Pfizer Inc., Amgen Inc., AbbVie Inc., Novartis, Johnson and Johnson, Mylan and Bayer.
Increase in incidence of autoimmune disease, surge in adoption of autoimmune disease therapeutics, and growth in R&D activities to develop ideal therapeutics are the key drivers of the global autoimmune disease therapeutics market. Furthermore, technological advancements in screening procedures, wide availability of therapeutics, and strong presence of pipeline drugs such as tocilizumab, baricitinib, certolizumab, secukinumab, etanercept, olokizumab, abatacept, apremilast, PF-06438179, golimumab, ustekinumab, etrolizumab, tofacitinib and others, are expected to further influence the market growth. On the other side, higher cost of advanced therapeutics can hamper the autoimmune disease therapeutics market growth.
Autoimmune Diseases Therapies Market Size & Market Analysis
The global autoimmune disease therapeutics market is segmented based on drug class, indication, sales channel, and region. Based on drug class, the market is classified as anti-inflammatory, antihyperglycemics, NSAIDs, interferons, and others. On the basis of indication rheumatic disease, type 1 diabetes, multiple sclerosis, inflammatory bowel disease, and other indications. According to the sales channel, the market is categorized as hospital pharmacy, drug stores & retail pharmacy, and online store. Based on region, the autoimmune disease therapeutics market size is studied across North America (U.S., Canada, and Mexico), Europe (Germany, France, the UK, Italy, Spain, and rest of Europe), Asia-Pacific (China, Japan, Australia, India, South Korea, and rest of Asia-Pacific), and LAMEA (Brazil, Saudi Arabia, and rest of LAMEA).
Based on drug class, the market is classified as anti-inflammatory, antihyperglycemics, NSAIDs, interferons, and others. Anti-inflammatory was the major revenue contributing segment in 2017 and is expected to exhibit dominance during the forecast period owing to increase in demand for anti- inflammatory drugs, launch of newer anti- inflammatory drugs with the supportive reimbursement policies for use of advanced anti- inflammatory drugs in some countries. Moreover, expected launch of pipeline anti-inflammatories such as tocilizumab, certolizumab, secukinumab, etanercept, golimumab and others, is estimated to further influence the market growth.
Anti-inflammatory drugs are preferably used in the treatment of inflammatory bowel disease, osteoarthritis, rheumatoid arthritis, fibromyalgia, systemic lupus erythematosus, gout, juvenile idiopathic arthritis, infectious arthritis, psoriatic arthritis, ankylosing spondylitis, reactive arthritis, scleroderma, polymyalgia rheumatica etc. Rise in incidence of these disease further fuels the market growth.
Presently, Adalimumab, Etanercept, Infliximab, Abatacept, Certolizumab, Golimumab, Ustekinumab, Tocilizumab, Secukinumab and others are widely used in the treatment of different form of autoimmune disease. Wide availability of anti-inflammatory drugs, preferable use of these drugs with favorable reimbursement in some countries contribute to the market growth.
Autoimmune Diseases Therapies Market Size & Market Analysis
Source : https://www.medgadget.com/2021/02/autoimmune-disorder-therapies-global-market-expected-to-reach-90-billion-by-2024.html
Source : https://www.alliedmarketresearch.com/autoimmune-disease-therapeutics-market